ITS Plot Examples: Univariate vs Multivariate, Two vs Three Periods#
This notebook illustrates the different plot options for multiple outcomes Interrupted Time Series (ITS) in CausalPy:
Two-period design (permanent intervention: one treatment start) vs three-period design (temporary intervention: treatment start and end)
For multivariate: overlay (all outcomes on the same axes) vs per_outcome (one figure per outcome)
All examples use the Bayesian (PyMC) workflow. Sampling uses reduced chains/draws so the notebook runs quickly.
Summary: What each part does#
1. Multivariate, two-period : Several outcomes at once, each with its own formula, using
MultivarLinearReg. Demonstrates two plot layouts: overlay (all outcomes on the same axes) and per_outcome (one figure per outcome, each with three panels).2. Multivariate, three-period : Same multivariate setup as above, but with a temporary intervention (start and end). Again shows overlay and per_outcome options.
3. Get plot data (DataFrame export) : How to obtain the data behind the plots via
get_plot_data(): for univariate ITS you get one DataFrame with time index, prediction, impact, and HDI columns; for multivariate you get a long-format DataFrame with anoutcomecolumn.4. Proving we recover the covariance matrix : Validation of
MultivarLinearReg: we generate data from a known residual covariance (three outcomes), fit the model, and compare the posterior mean covariance to the true one (max absolute difference and mean squared error).7. Summary : How
summary()works: it prints the formula, coefficients, and the effect summary table (per outcome in the multivariate case).
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import causalpy as cp
seed = 42
# Fast sampling for demo (reduce for real analyses)
sample_kwargs = {
"chains": 2,
"draws": 200,
"tune": 100,
"progressbar": False,
"random_seed": seed,
}
import os
os.environ["KMP_DUPLICATE_LIB_OK"] = "TRUE"
1. Multivariate, two-period#
Two outcomes with a list of formulas. We use MultivarLinearReg (supports multivariate). Plot API:
Default: call
plot()(orplot(layout="overlay")) — one figure, 3 panels, all outcomes on the same axes.Per outcome: call
plot(layout="per_outcome")— one figure per outcome, each with 3 panels.Optionally pass
outcomes_to_plot=["y1", ...]to show a subset of outcomes.
rng = np.random.default_rng(seed)
n = 72
dates = pd.date_range(start="2015-01-01", periods=n, freq="MS")
t = np.arange(n, dtype=float)
y1 = 10 + 0.3 * t + 2 * np.sin(2 * np.pi * np.arange(n) / 12) + rng.normal(0, 1, n)
y2 = 20 + 0.1 * t + rng.normal(0, 1.5, n)
df_multi = pd.DataFrame(
{"y1": y1, "y2": y2, "t": t},
index=dates,
)
df_multi.index.name = "obs_ind"
treatment_time_m = pd.to_datetime("2018-01-01")
result_multi_2p = cp.InterruptedTimeSeries(
df_multi,
treatment_time_m,
formula=["y1 ~ 1 + t", "y2 ~ 1 + t"],
model=cp.pymc_models.MultivarLinearReg(sample_kwargs=sample_kwargs),
)
c:\Users\jeanv\miniconda3\envs\CausalPy\Lib\site-packages\threadpoolctl.py:1226: RuntimeWarning:
Found Intel OpenMP ('libiomp') and LLVM OpenMP ('libomp') loaded at
the same time. Both libraries are known to be incompatible and this
can cause random crashes or deadlocks on Linux when loaded in the
same Python program.
Using threadpoolctl may cause crashes or deadlocks. For more
information and possible workarounds, please see
https://github.com/joblib/threadpoolctl/blob/master/multiple_openmp.md
warnings.warn(msg, RuntimeWarning)
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [beta, chol_cov]
Sampling 2 chains for 100 tune and 200 draw iterations (200 + 400 draws total) took 35 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Sampling: [beta, chol_cov, y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
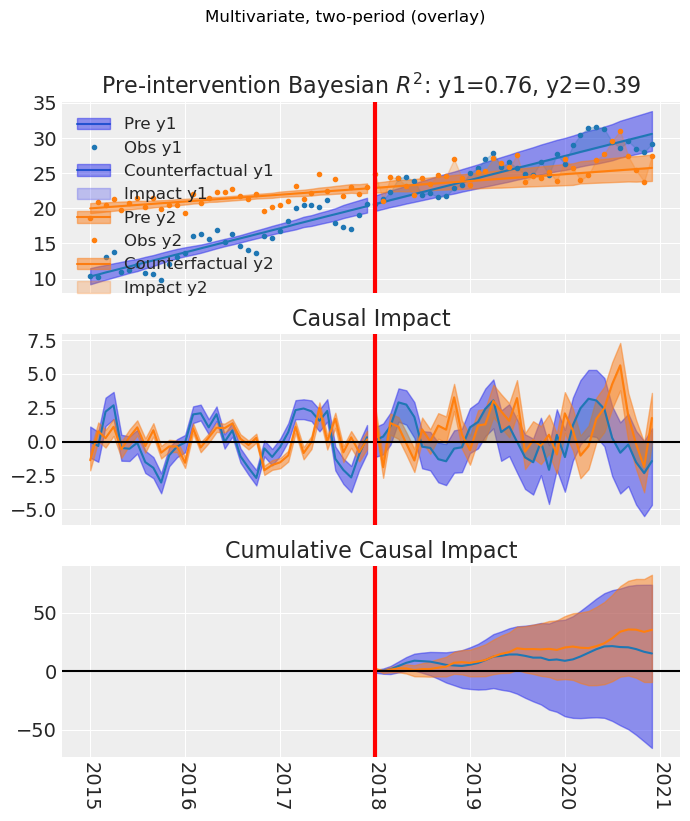
# Default: overlay (all outcomes on the same axes). Same as plot(layout="overlay")
fig, ax = result_multi_2p.plot()
fig.suptitle("Multivariate, two-period (overlay)", y=1.02)
fig.tight_layout()
plt.show()
C:\Users\jeanv\AppData\Local\Temp\ipykernel_12588\2607712260.py:4: UserWarning: The figure layout has changed to tight
fig.tight_layout()
Inferred covariance matrix#
MultivarLinearReg estimates a covariance matrix between outcomes (LKJ prior on the correlation and scale per outcome). Below we inspect the posterior mean covariance and the derived correlation matrix to show that the model correctly infers the residual dependence between y1 and y2.
# Layout per_outcome: one figure per outcome (returns a list of (fig, ax))
# One figure per outcome: pass layout="per_outcome"
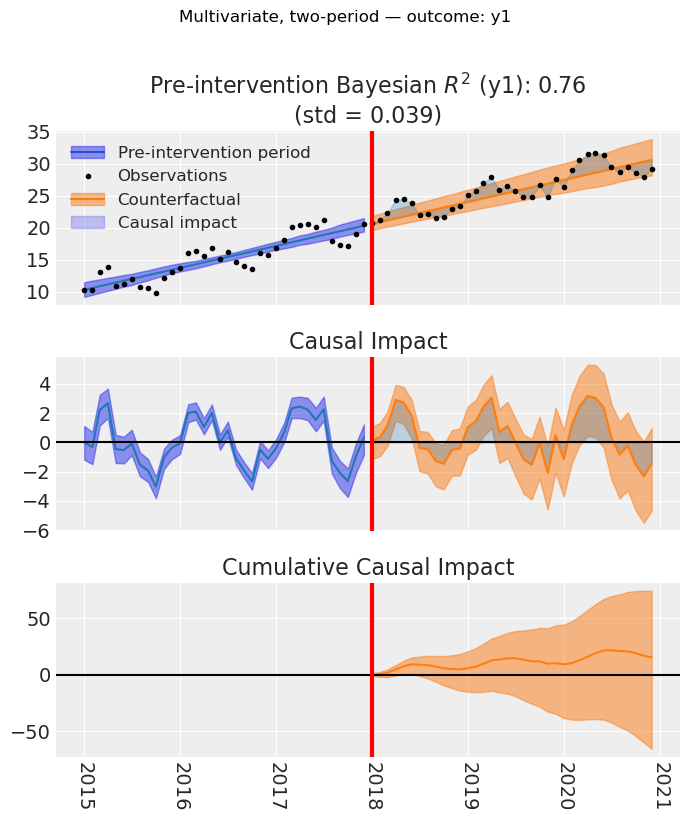
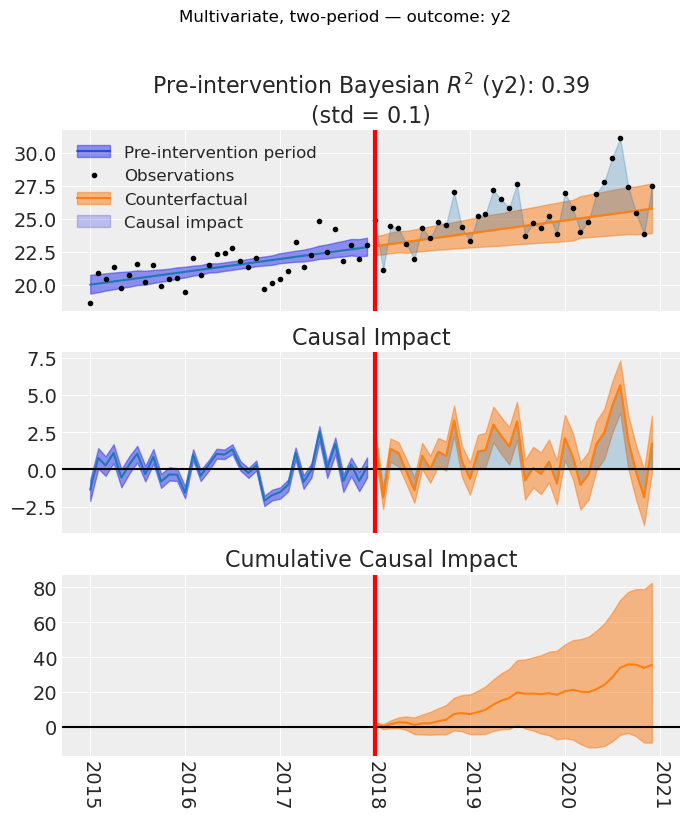
figs = result_multi_2p.plot(layout="per_outcome")
for i, (fig, _ax) in enumerate(figs):
fig.suptitle(
f"Multivariate, two-period — outcome: {result_multi_2p.outcome_variable_names[i]}",
y=1.02,
)
fig.tight_layout()
plt.show()
C:\Users\jeanv\AppData\Local\Temp\ipykernel_7320\885547691.py:9: UserWarning: The figure layout has changed to tight
fig.tight_layout()
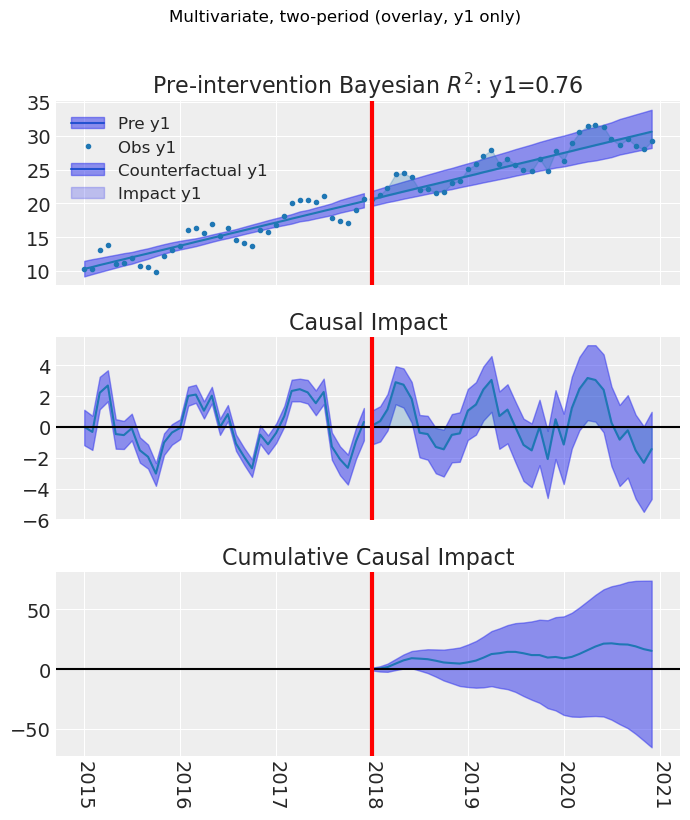
# Optional: subset of outcomes (overlay). Default overlay + outcomes_to_plot
fig, ax = result_multi_2p.plot(outcomes_to_plot=["y1"])
fig.suptitle("Multivariate, two-period (overlay, y1 only)", y=1.02)
fig.tight_layout()
plt.show()
C:\Users\jeanv\AppData\Local\Temp\ipykernel_14100\1352864922.py:4: UserWarning: The figure layout has changed to tight
fig.tight_layout()
4. Multivariate, three-period#
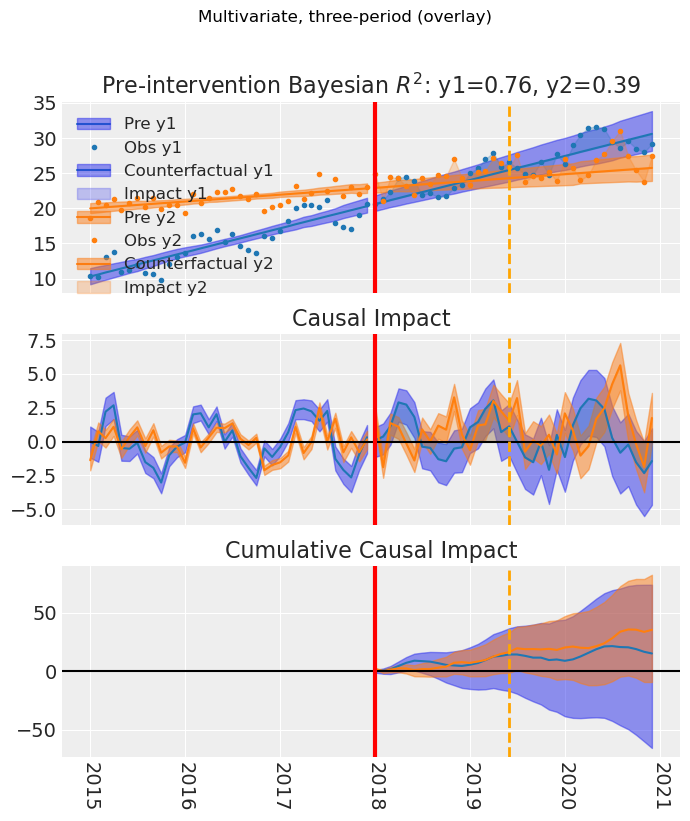
Multivariate ITS with temporary intervention: same overlay and per_outcome options, with treatment start and end lines on the plot.
treatment_end_time_m = pd.to_datetime("2019-06-01")
result_multi_3p = cp.InterruptedTimeSeries(
df_multi,
treatment_time_m,
formula=["y1 ~ 1 + t", "y2 ~ 1 + t"],
treatment_end_time=treatment_end_time_m,
model=cp.pymc_models.MultivarLinearReg(sample_kwargs=sample_kwargs),
)
fig, ax = result_multi_3p.plot() # default overlay
fig.suptitle("Multivariate, three-period (overlay)", y=1.02)
fig.tight_layout()
plt.show()
c:\Users\jeanv\miniconda3\envs\CausalPy\Lib\site-packages\threadpoolctl.py:1226: RuntimeWarning:
Found Intel OpenMP ('libiomp') and LLVM OpenMP ('libomp') loaded at
the same time. Both libraries are known to be incompatible and this
can cause random crashes or deadlocks on Linux when loaded in the
same Python program.
Using threadpoolctl may cause crashes or deadlocks. For more
information and possible workarounds, please see
https://github.com/joblib/threadpoolctl/blob/master/multiple_openmp.md
warnings.warn(msg, RuntimeWarning)
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [beta, chol_cov]
Sampling 2 chains for 100 tune and 200 draw iterations (200 + 400 draws total) took 18 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Sampling: [beta, chol_cov, y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
C:\Users\jeanv\AppData\Local\Temp\ipykernel_14100\1244863932.py:12: UserWarning: The figure layout has changed to tight
fig.tight_layout()
5. Get plot data (DataFrame export)#
For univariate ITS, get_plot_data() returns a DataFrame with the time index, prediction, impact, and HDI columns. For multivariate, the DataFrame is in long format with an outcome column.
# Multivariate: long format with outcome column
plot_data_multi = result_multi_2p.get_plot_data()
print("Multivariate columns:", list(plot_data_multi.columns))
print("Outcomes in data:", plot_data_multi["outcome"].unique().tolist())
plot_data_multi.head(10)
Multivariate columns: ['y1', 'y2', 't', 'outcome', 'prediction', 'pred_hdi_lower_94', 'pred_hdi_upper_94', 'impact', 'impact_hdi_lower_94', 'impact_hdi_upper_94']
Outcomes in data: ['y1', 'y2']
| y1 | y2 | t | outcome | prediction | pred_hdi_lower_94 | pred_hdi_upper_94 | impact | impact_hdi_lower_94 | impact_hdi_upper_94 | |
|---|---|---|---|---|---|---|---|---|---|---|
| obs_ind | ||||||||||
| 2015-01-01 | 10.304717 | 18.620822 | 0.0 | y1 | 10.298209 | 9.191484 | 11.490822 | 0.006508 | -1.065639 | 1.347100 |
| 2015-02-01 | 10.260016 | 20.845741 | 1.0 | y1 | 10.584169 | 9.539022 | 11.750634 | -0.324153 | -1.354346 | 0.943285 |
| 2015-03-01 | 13.082502 | 20.413639 | 2.0 | y1 | 10.870129 | 9.836503 | 11.938577 | 2.212373 | 1.224135 | 3.403732 |
| 2015-04-01 | 13.840565 | 21.335728 | 3.0 | y1 | 11.156089 | 10.167732 | 12.176982 | 2.684475 | 1.738196 | 3.799754 |
| 2015-05-01 | 10.981016 | 19.759121 | 4.0 | y1 | 11.442050 | 10.477895 | 12.392716 | -0.461034 | -1.364177 | 0.578165 |
| 2015-06-01 | 11.197820 | 20.737810 | 5.0 | y1 | 11.728010 | 10.809258 | 12.628730 | -0.530189 | -1.391565 | 0.473865 |
| 2015-07-01 | 11.927840 | 21.538386 | 6.0 | y1 | 12.013970 | 11.068768 | 12.802749 | -0.086130 | -0.905423 | 0.843753 |
| 2015-08-01 | 10.783757 | 20.235980 | 7.0 | y1 | 12.299930 | 11.457413 | 13.099959 | -1.516173 | -2.284106 | -0.637511 |
| 2015-09-01 | 10.651148 | 21.485163 | 8.0 | y1 | 12.585890 | 11.794811 | 13.352568 | -1.934742 | -2.670125 | -1.089803 |
| 2015-10-01 | 9.846956 | 19.907111 | 9.0 | y1 | 12.871851 | 12.189518 | 13.664122 | -3.024894 | -3.736210 | -2.232080 |
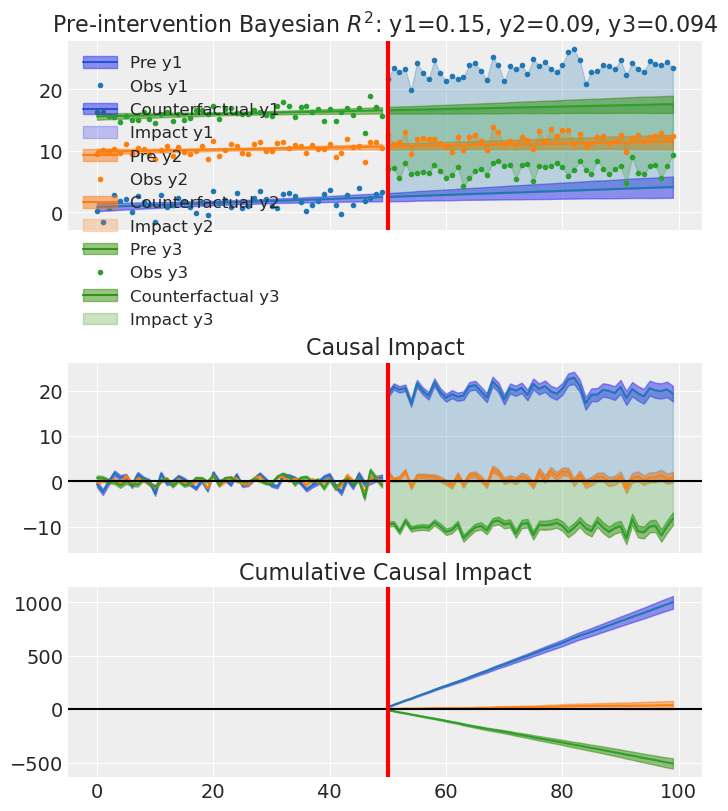
6. Proving we recover the covariance matrix#
To show that MultivarLinearReg correctly infers the residual covariance between outcomes, we generate a dataset with three outcomes whose residuals follow a known covariance matrix. We then fit the model and compare the posterior mean covariance to the true one. For this section we use heavier sampling (more chains, draws, tune) and target_accept=0.95 so that NUTS takes smaller steps;
# True covariance matrix (3x3, positive definite) used to generate correlated residuals
# We choose a known structure: correlation + scale per outcome
true_corr = np.array(
[
[1.0, 0.6, 0.2],
[0.6, 1.0, 0.5],
[0.2, 0.5, 1.0],
]
)
true_std = np.array([1.2, 0.9, 1.0]) # standard deviations per outcome
true_cov = np.outer(true_std, true_std) * true_corr
outcome_names = ["y1", "y2", "y3"]
# Generate data: trend + correlated noise
n = 100
rng_cov = np.random.default_rng(seed)
t = np.linspace(0, 10, n)
# Mean structure: different intercept and slope per outcome
beta_true = np.array(
[
[1.0, 0.3], # y1: intercept, slope
[10.0, 0.1], # y2
[15.5, 0.2], # y3
]
)
X = np.column_stack([np.ones(n), t])
mean_3d = X @ beta_true.T # (n, 3)
# Correlated residuals from the known covariance; standardize so residual variance = 1 per outcome
# (reduces sampler geometry issues and we then compare inferred cov to true_corr)
residuals = rng_cov.multivariate_normal(np.zeros(3), true_cov, size=n)
residuals_std = residuals / residuals.std(axis=0) * true_std
y1 = mean_3d[:, 0] + residuals_std[:, 0]
y2 = mean_3d[:, 1] + residuals_std[:, 1]
y3 = mean_3d[:, 2] + residuals_std[:, 2]
# Treatment effect from t=50 onward
treatment_idx = 50
y1[treatment_idx:] += 20.0
y2[treatment_idx:] += 1.0
y3[treatment_idx:] -= 10
df_cov = pd.DataFrame(
{"y1": y1, "y2": y2, "y3": y3, "t": t},
index=np.arange(n),
)
df_cov.index.name = "obs_ind"
treatment_time_cov = 50.0 # corresponds to index ~50
# Heavier sampling + target_accept to avoid divergences (LKJ covariance has difficult geometry)
sample_kwargs_cov = {
"chains": 4,
"draws": 4000,
"tune": 2000,
"target_accept": 0.98,
"progressbar": True,
"random_seed": seed,
}
result_cov = cp.InterruptedTimeSeries(
df_cov,
treatment_time=treatment_time_cov,
formula=["y1 ~ 1 + t", "y2 ~ 1 + t", "y3 ~ 1 + t"],
model=cp.pymc_models.MultivarLinearReg(sample_kwargs=sample_kwargs_cov),
)
c:\Users\jeanv\miniconda3\envs\CausalPy\Lib\site-packages\threadpoolctl.py:1226: RuntimeWarning:
Found Intel OpenMP ('libiomp') and LLVM OpenMP ('libomp') loaded at
the same time. Both libraries are known to be incompatible and this
can cause random crashes or deadlocks on Linux when loaded in the
same Python program.
Using threadpoolctl may cause crashes or deadlocks. For more
information and possible workarounds, please see
https://github.com/joblib/threadpoolctl/blob/master/multiple_openmp.md
warnings.warn(msg, RuntimeWarning)
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [beta, chol_cov]
Sampling 4 chains for 2_000 tune and 4_000 draw iterations (8_000 + 16_000 draws total) took 135 seconds.
Sampling: [beta, chol_cov, y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
result_cov.plot() # default overlay
plt.show()
# Compare true vs inferred covariance matrix
idata_cov = result_cov.model.idata
cov_post = idata_cov.posterior["cov"].mean(dim=["chain", "draw"]).values
if cov_post.ndim > 2:
cov_post = cov_post.reshape(cov_post.shape[-2], cov_post.shape[-1])
# True residual cov = true_corr (we used standardized residuals, so unit variance)
true_cov_df = pd.DataFrame(true_cov, index=outcome_names, columns=outcome_names)
inferred_cov_df = pd.DataFrame(cov_post, index=outcome_names, columns=outcome_names)
diff_cov = inferred_cov_df - true_cov_df
print("True residual covariance (correlation matrix, from standardized residuals):")
display(true_cov_df)
print("Inferred posterior mean covariance matrix:")
display(inferred_cov_df)
print("Difference (inferred - true). Should be close to zero:")
display(diff_cov.round(4))
print("Max absolute difference: {:.4f}".format(np.abs(diff_cov.values).max()))
print("Mean squared error: {:.6f}".format(np.mean(diff_cov.values**2)))
True residual covariance (correlation matrix, from standardized residuals):
| y1 | y2 | y3 | |
|---|---|---|---|
| y1 | 1.440 | 0.648 | 0.24 |
| y2 | 0.648 | 0.810 | 0.45 |
| y3 | 0.240 | 0.450 | 1.00 |
Inferred posterior mean covariance matrix:
| y1 | y2 | y3 | |
|---|---|---|---|
| y1 | 1.551499 | 0.514865 | 0.086018 |
| y2 | 0.514865 | 0.697577 | 0.539261 |
| y3 | 0.086018 | 0.539261 | 1.016035 |
Difference (inferred - true). Should be close to zero:
| y1 | y2 | y3 | |
|---|---|---|---|
| y1 | 0.1115 | -0.1331 | -0.1540 |
| y2 | -0.1331 | -0.1124 | 0.0893 |
| y3 | -0.1540 | 0.0893 | 0.0160 |
Max absolute difference: 0.1540
Mean squared error: 0.013793
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# 1. Extract all raw posterior samples for the 'cov' matrix
# The initial shape is usually (chains, draws, N, N)
cov_samples = idata_cov.posterior["cov"].values
# Flatten the chain and draw dimensions to pool all samples together
# New shape: (total_samples, N, N)
cov_samples_flat = cov_samples.reshape(-1, cov_samples.shape[-2], cov_samples.shape[-1])
n_outcomes = len(outcome_names)
# 2. Set up a grid of subplots (N x N)
fig, axes = plt.subplots(
n_outcomes, n_outcomes, figsize=(3.5 * n_outcomes, 3.5 * n_outcomes)
)
# Handle the 1x1 case gracefully if there's only one outcome
if n_outcomes == 1:
axes = np.array([[axes]])
# 3. Loop through each cell in the covariance matrix
for i in range(n_outcomes):
for j in range(n_outcomes):
ax = axes[i, j]
# Extract the 1D array of samples for this specific (i, j) component
cell_samples = cov_samples_flat[:, i, j]
# Plot the posterior distribution
sns.kdeplot(cell_samples, ax=ax, fill=True, color="C0", alpha=0.6)
# Plot the true value as a vertical red dashed line
true_val = true_cov[i, j]
ax.axvline(
true_val,
color="red",
linestyle="--",
linewidth=2,
label=f"True: {true_val:.2f}",
)
# Plot the inferred mean as a vertical black dotted line
inferred_mean = cov_post[i, j]
ax.axvline(
inferred_mean,
color="black",
linestyle=":",
linewidth=2,
label=f"Mean: {inferred_mean:.2f}",
)
# Formatting
ax.set_title(f"Cov({outcome_names[i]}, {outcome_names[j]})")
ax.set_yticks([]) # Hide y-axis ticks for cleaner look
# Only show the legend on the very first subplot to avoid clutter
if i == 0 and j == 0:
ax.legend()
plt.tight_layout()
plt.show()
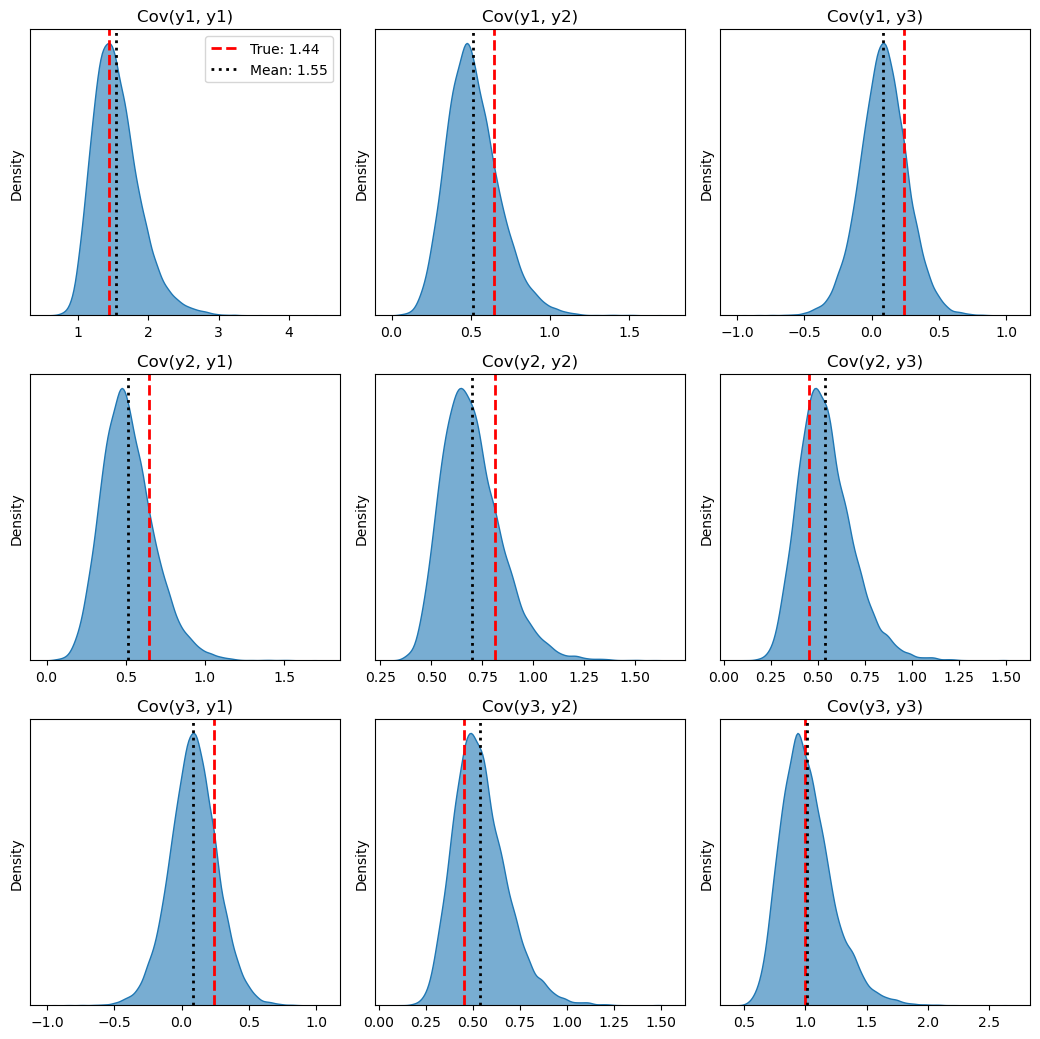
7. Summary: coefficients and effect summary#
The summary() method prints the experiment type, formula, model coefficients, and an effect summary table (mean impact, HDI, cumulative effect, etc.). For multivariate ITS, the effect summary has one row per outcome and per statistic (average / cumulative), so you can read the causal effect for each outcome.
# Uses the multivariate two-period result from section 1
result_cov.summary()
==================================Pre-Post Fit==================================
Formula: ['y1 ~ 1 + t', 'y2 ~ 1 + t', 'y3 ~ 1 + t']
Model coefficients:
Outcome: y1
Intercept 0.86, 94% HDI [0.21, 1.5]
t 0.33, 94% HDI [0.1, 0.55]
Outcome: y2
Intercept 9.9, 94% HDI [9.4, 10]
t 0.16, 94% HDI [0.0099, 0.31]
Outcome: y3
Intercept 16, 94% HDI [15, 16]
t 0.2, 94% HDI [0.017, 0.38]
Residual covariance (upper triangle), 94% HDI:
cov(y1, y1) 1.6, 94% HDI [1, 2.3]
cov(y1, y2) 0.51, 94% HDI [0.24, 0.87]
cov(y1, y3) 0.086, 94% HDI [-0.26, 0.43]
cov(y2, y2) 0.7, 94% HDI [0.48, 1]
cov(y2, y3) 0.54, 94% HDI [0.31, 0.85]
cov(y3, y3) 1, 94% HDI [0.69, 1.5]
Effect summary:
outcome statistic mean median hdi_lower hdi_upper p_gt_0 relative_mean relative_hdi_lower relative_hdi_upper
0 y1 average 20.038780 20.036233 18.789462 21.273351 1.000000 634.875454 369.626710 936.991426
1 y1 cumulative 1001.938982 1001.811646 939.473116 1063.667528 1.000000 634.875456 369.626711 936.991430
2 y2 average 0.750040 0.748689 -0.075078 1.576197 0.962562 6.912498 -0.923358 14.976382
3 y2 cumulative 37.502004 37.434429 -3.753891 78.809834 0.962562 6.912498 -0.923358 14.976382
4 y3 average -10.165110 -10.166098 -11.158164 -9.155131 0.000000 -59.417583 -61.792658 -57.032796
5 y3 cumulative -508.255516 -508.304923 -557.908203 -457.756544 0.000000 -59.417583 -61.792658 -57.032796
[y1] Post-period (50 to 99), the average effect was 20.04 (95% HDI [18.79, 21.27]), with a posterior probability of an increase of 1.000. The cumulative effect was 1001.94 (95% HDI [939.47, 1063.67]); probability of an increase 1.000. Relative to the counterfactual, this equals 634.88% on average (95% HDI [369.63%, 936.99%]).
[y2] Post-period (50 to 99), the average effect was 0.75 (95% HDI [-0.08, 1.58]), with a posterior probability of an increase of 0.963. The cumulative effect was 37.50 (95% HDI [-3.75, 78.81]); probability of an increase 0.963. Relative to the counterfactual, this equals 6.91% on average (95% HDI [-0.92%, 14.98%]).
[y3] Post-period (50 to 99), the average effect was -10.17 (95% HDI [-11.16, -9.16]), with a posterior probability of an increase of 0.000. The cumulative effect was -508.26 (95% HDI [-557.91, -457.76]); probability of an increase 0.000. Relative to the counterfactual, this equals -59.42% on average (95% HDI [-61.79%, -57.03%]).

